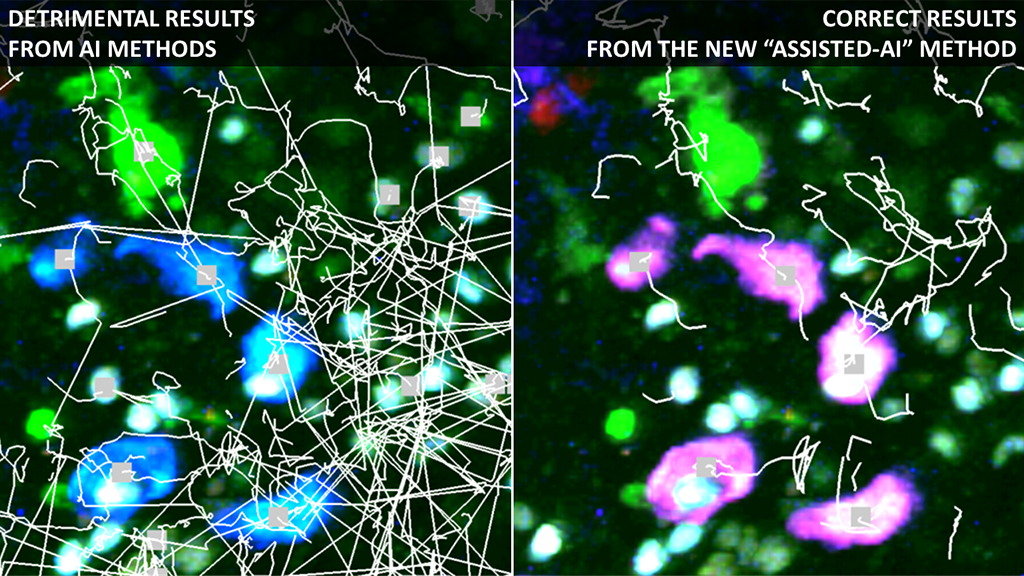
CANCOL (Imaris plugin & standalone)
Ccreates an additional virtual imaging channel (magenta) specific for the cells of interest.
Enhances the automatic tracking of immune cells, avoiding mistakes possibly introduced by AI.
--> Useful when cells emit in multipe acquisition channels or when they become dim.
The user just has to draw a few lines on the cells of interest and other lines on the background or other cells.
- Imaris XTension and standalone version (download)
- Article at The Journal of Immunology https://doi.org/10.4049/jimmunol.2100811
- Preprint (with preliminary results, final paper is different): https://www.biorxiv.org/content/10.1101/829838v2
Example results:
Motility heatmap
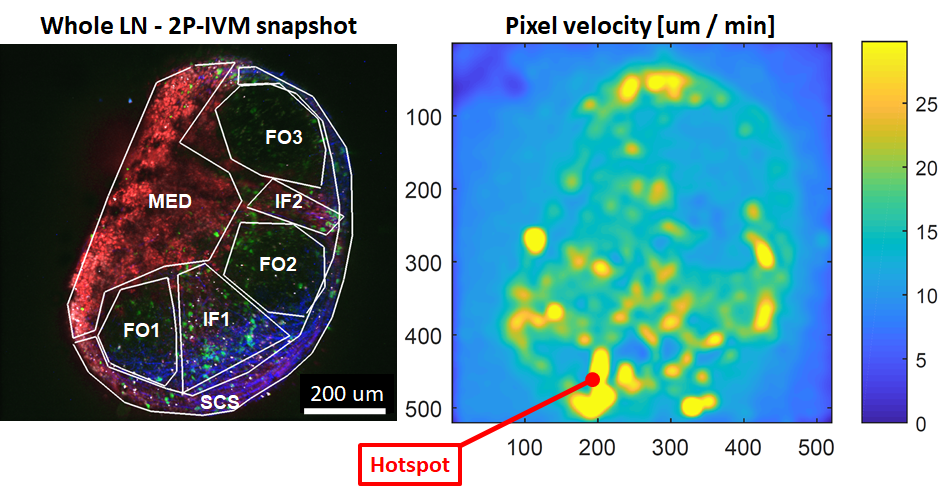
This tool reads Imaris files (ims) and creates a motility heatmap based on Optical Flow as described in Pizzagalli et al. Frontiers in Immunology (2019)
It is useful to identify hotspots with high motility, or to analyze cell motility without requiring tracking.
- Standalone program for Windows, Imaris plugins, and Open Source code available here
- Please cite as Pizzagalli DU & Latino I et al., Frontiers in immunology (2019) https://www.frontiersin.org/articles/10.3389/fimmu.2019.02621/abstract
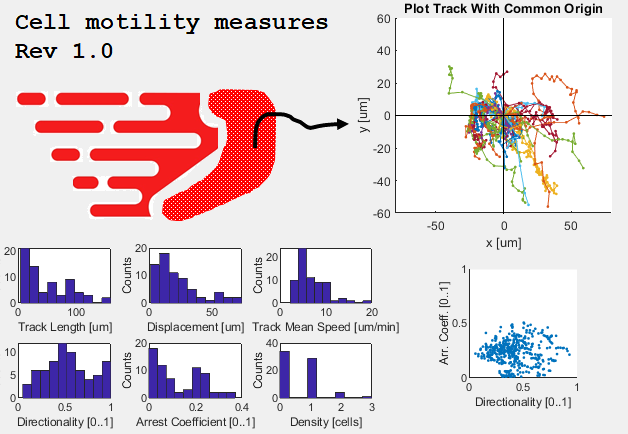
General motility measures (track based and step based)
This tool reads Imaris exported xls files, as well as TrackMate xml files and computes basic motility patterns useful in intravital imaging studies including:
track length, speed, directionality, displacement, arrest coefficient, density. Moroever, it creates plots of track with common origin.
download link will be provided before the aiivm workshop of May2022
Cell action recognition (track-based)
This tool to detects and count actions of neutrophils as described in Pizzagalli et al. Frontiers in Immunology (2019)
https://www.frontiersin.org/articles/10.3389/fimmu.2019.02621/abstract
Like a FACS gating, but for motility patterns
download link will be provided before the aiivm workshop of May2022